---
status: w3c/CG-FINAL
level: 1
date: 2023-05-25
version: 0.4
title: Next-generation file formats (NGFF)
abstract: >-
This document contains next-generation file format (NGFF)
specifications for storing bioimaging data in the cloud.
All specifications are submitted to the https://image.sc community for review.
authors:
- name: "Editor: Josh Moore"
roles: Editor
affiliations:
- name: University of Dundee (UoD)
ror: https://ror.org/03h2bxq36
url: https://www.dundee.ac.uk
- name: "Editor: Sébastien Besson"
roles: Editor
orcid: https://orcid.org/0000-0001-8783-1429
affiliations:
- name: University of Dundee (UoD)
ror: https://ror.org/03h2bxq36
url: https://www.dundee.ac.uk
- name: "Editor: Constantin Pape"
roles: Editor
orcid: 0000-0001-6562-7187
affiliations:
- name: European Molecular Biology Laboratory (EMBL)
url: https://www.embl.org/sites/heidelberg/
ror: https://ror.org/03mstc592
---
# Version 0.4
## Introduction
(version0.4:intro)=
Bioimaging science is at a crossroads. Currently, the drive to acquire more,
larger, preciser spatial measurements is unfortunately at odds with our ability
to structure and share those measurements with others. During a global pandemic
more than ever, we believe fervently that global, collaborative discovery as
opposed to the post-publication, "data-on-request" mode of operation is the
path forward. Bioimaging data should be shareable via open and commercial cloud
resources without the need to download entire datasets.
At the moment, that is not the norm. The plethora of data formats produced by
imaging systems are ill-suited to remote sharing. Individual scientists
typically lack the infrastructure they need to host these data themselves. When
they acquire images from elsewhere, time-consuming translations and data
cleaning are needed to interpret findings. Those same costs are multiplied when
gathering data into online repositories where curator time can be the limiting
factor before publication is possible. Without a common effort, each lab or
resource is left building the tools they need and maintaining that
infrastructure often without dedicated funding.
This document defines a specification for bioimaging data to make it possible
to enable the conversion of proprietary formats into a common, cloud-ready one.
Such next-generation file formats layout data so that individual portions, or
"chunks", of large data are reference-able eliminating the need to download
entire datasets.
### Why "NGFF"?
(version0.4:why-ngff)=
A short description of what is needed for an imaging format is "a hierarchy
of n-dimensional (dense) arrays with metadata". This combination of features
is certainly provided by HDF5
from the HDF Group, which a number of
bioimaging formats do use. HDF5 and other larger binary structures, however,
are ill-suited for storage in the cloud where accessing individual chunks
of data by name rather than seeking through a large file is at the heart of
parallelization.
As a result, a number of formats have been developed more recently which provide
the basic data structure of an HDF5 file, but do so in a more cloud-friendly way.
In the [PyData](https://pydata.org/) community, the Zarr ({cite:t}`zarr`) format was developed
for easily storing collections of [NumPy](https://numpy.org/) arrays. In the
[ImageJ](https://imagej.net/) community, N5 ({cite:t}`n5`) was developed to work around
the limitations of HDF5 ("N5" was originally short for "Not-HDF5").
Both of these formats permit storing individual chunks of data either locally in
separate files or in cloud-based object stores as separate keys.
A [current effort](https://zarr-specs.readthedocs.io/en/core-protocol-v3.0-dev/protocol/core/v3.0.html)
is underway to unify the two similar specifications to provide a single binary
specification. The editor's draft will soon be entering a [request for comments (RFC)](https://github.com/zarr-developers/zarr-specs/issues/101) phase with the goal of having a first version early in 2021. As that
process comes to an end, this document will be updated.
### OME-NGFF
(version0.4:ome-ngff)=
The conventions and specifications defined in this document are designed to
enable next-generation file formats to represent the same bioimaging data
that can be represented in \[OME-TIFF](http://www.openmicroscopy.org/ome-files/)
and beyond. However, the conventions will also be usable by HDF5 and other sufficiently advanced
binary containers. Eventually, we hope, the moniker "next-generation" will no longer be
applicable, and this will simply be the most efficient, common, and useful representation
of bioimaging data, whether during acquisition or sharing in the cloud.
Note: The following text makes use of OME-Zarr ({cite:t}`ome-zarr-py`), the current prototype implementation,
for all examples.
### Document conventions
The key words “MUST”, “MUST NOT”, “REQUIRED”, “SHALL”, “SHALL NOT”, “SHOULD”, “SHOULD NOT”,
“RECOMMENDED”, “MAY”, and “OPTIONAL” are to be interpreted as described in
[RFC 2119](https://tools.ietf.org/html/rfc2119).
Transitional metadata is added to the specification with the
intention of removing it in the future. Implementations may be expected (MUST) or
encouraged (SHOULD) to support the reading of the data, but writing will usually
be optional (MAY). Examples of transitional metadata include custom additions by
implementations that are later submitted as a formal specification. (See (bioformats2raw metadata)(#version0.4:bf2raw))
Some of the JSON examples in this document include comments. However, these are only for
clarity purposes and comments MUST NOT be included in JSON objects.
## On-disk (or in-cloud) layout
(version0.4:on-disk)=
An overview of the layout of an OME-Zarr fileset should make
understanding the following metadata sections easier. The hierarchy
is represented here as it would appear locally but could equally
be stored on a web server to be accessed via HTTP or in object storage
like S3 or GCS.
OME-Zarr is an implementation of the OME-NGFF specification using the Zarr
format. Arrays MUST be defined and stored in a hierarchical organization as
defined by the
[version 2 of the Zarr specification ](https://zarr.readthedocs.io/en/stable/spec/v2.html).
OME-NGFF metadata MUST be stored as attributes in the corresponding Zarr
groups.
### Images
(version0.4:image-layout)=
The following layout describes the expected Zarr hierarchy for images with
multiple levels of resolutions and optionally associated labels.
Note that the number of dimensions is variable between 2 and 5 and that axis names are arbitrary, see (multiscales metadata)(#version0.4:multiscale-md) for details.
For this example we assume an image with 5 dimensions and axes called `t,c,z,y,x`.
. # Root folder, potentially in S3,
│ # with a flat list of images by image ID.
│
├── 123.zarr # One image (id=123) converted to Zarr.
│
└── 456.zarr # Another image (id=456) converted to Zarr.
│
├── .zgroup # Each image is a Zarr group, or a folder, of other groups and arrays.
├── .zattrs # Group level attributes are stored in the .zattrs file and include
│ # "multiscales" and "omero" (see below). In addition, the group level attributes
│ # may also contain "_ARRAY_DIMENSIONS" for compatibility with xarray if this group directly contains multi-scale arrays.
│
├── 0 # Each multiscale level is stored as a separate Zarr array,
│ ... # which is a folder containing chunk files which compose the array.
├── n # The name of the array is arbitrary with the ordering defined by
│ │ # by the "multiscales" metadata, but is often a sequence starting at 0.
│ │
│ ├── .zarray # All image arrays must be up to 5-dimensional
│ │ # with the axis of type time before type channel, before spatial axes.
│ │
│ └─ t # Chunks are stored with the nested directory layout.
│ └─ c # All but the last chunk element are stored as directories.
│ └─ z # The terminal chunk is a file. Together the directory and file names
│ └─ y # provide the "chunk coordinate" (t, c, z, y, x), where the maximum coordinate
│ └─ x # will be `dimension_size / chunk_size`.
│
└── labels
│
├── .zgroup # The labels group is a container which holds a list of labels to make the objects easily discoverable
│
├── .zattrs # All labels will be listed in `.zattrs` e.g. `{ "labels": [ "original/0" ] }`
│ # Each dimension of the label `(t, c, z, y, x)` should be either the same as the
│ # corresponding dimension of the image, or `1` if that dimension of the label
│ # is irrelevant.
│
└── original # Intermediate folders are permitted but not necessary and currently contain no extra metadata.
│
└── 0 # Multiscale, labeled image. The name is unimportant but is registered in the "labels" group above.
├── .zgroup # Zarr Group which is both a multiscaled image as well as a labeled image.
├── .zattrs # Metadata of the related image and as well as display information under the "image-label" key.
│
├── 0 # Each multiscale level is stored as a separate Zarr array, as above, but only integer values
│ ... # are supported.
└── n
### High-content screening
(version0.4:hcs-layout)=
The following specification defines the hierarchy for a high-content screening
dataset. Three groups MUST be defined above the images:
- the group above the images defines the well and MUST implement the
[well specification](#version0.4:well-md). All images contained in a well are fields
of view of the same well
- the group above the well defines a row of wells
- the group above the well row defines an entire plate i.e. a two-dimensional
collection of wells organized in rows and columns. It MUST implement the
[plate specification](#version0.4:plate-md)
A well row group SHOULD NOT be present if there are no images in the well row.
A well group SHOULD NOT be present if there are no images in the well.
. # Root folder, potentially in S3,
│
└── 5966.zarr # One plate (id=5966) converted to Zarr
├── .zgroup
├── .zattrs # Implements "plate" specification
├── A # First row of the plate
│ ├── .zgroup
│ │
│ ├── 1 # First column of row A
│ │ ├── .zgroup
│ │ ├── .zattrs # Implements "well" specification
│ │ │
│ │ ├── 0 # First field of view of well A1
│ │ │ │
│ │ │ ├── .zgroup
│ │ │ ├── .zattrs # Implements "multiscales", "omero"
│ │ │ ├── 0
│ │ │ │ ... # Resolution levels
│ │ │ ├── n
│ │ │ └── labels # Labels (optional)
│ │ ├── ... # Fields of view
│ │ └── m
│ ├── ... # Columns
│ └── 12
├── ... # Rows
└── H
## Metadata
(version0.4:metadata)=
The various `.zattrs` files throughout the above array hierarchy may contain metadata
keys as specified below for discovering certain types of data, especially images.
### "axes" metadata
(version0.4:axes-md)
"axes" describes the dimensions of a physical coordinate space. It is a list of dictionaries, where each dictionary describes a dimension (axis) and:
- MUST contain the field "name" that gives the name for this dimension. The values MUST be unique across all "name" fields.
- SHOULD contain the field "type". It SHOULD be one of "space", "time" or "channel", but MAY take other string values for custom axis types that are not part of this specification yet.
- SHOULD contain the field "unit" to specify the physical unit of this dimension. The value SHOULD be one of the following strings, which are valid units according to UDUNITS-2.
- Units for "space" axes: 'angstrom', 'attometer', 'centimeter', 'decimeter', 'exameter', 'femtometer', 'foot', 'gigameter', 'hectometer', 'inch', 'kilometer', 'megameter', 'meter', 'micrometer', 'mile', 'millimeter', 'nanometer', 'parsec', 'petameter', 'picometer', 'terameter', 'yard', 'yoctometer', 'yottameter', 'zeptometer', 'zettameter'
- Units for "time" axes: 'attosecond', 'centisecond', 'day', 'decisecond', 'exasecond', 'femtosecond', 'gigasecond', 'hectosecond', 'hour', 'kilosecond', 'megasecond', 'microsecond', 'millisecond', 'minute', 'nanosecond', 'petasecond', 'picosecond', 'second', 'terasecond', 'yoctosecond', 'yottasecond', 'zeptosecond', 'zettasecond'
If part of (multiscales metadata)(#version0.4:multiscale-md), the length of "axes" MUST be equal to the number of dimensions of the arrays that contain the image data.
### "bioformats2raw.layout" (transitional)
(version0.4:bf2raw)=
[=Transitional=] "bioformats2raw.layout" metadata identifies a group which implicitly describes a series of images.
The need for the collection stems from the common "multi-image file" scenario in microscopy. Parsers like Bio-Formats
define a strict, stable ordering of the images in a single container that can be used to refer to them by other tools.
In order to capture that information within an OME-NGFF dataset, `bioformats2raw` internally introduced a wrapping layer.
The bioformats2raw layout has been added to v0.4 as a transitional specification to specify filesets that already exist
in the wild. An upcoming NGFF specification will replace this layout with explicit metadata.
#### Layout
(version0.4:bf2raw-layout)=
Typical Zarr layout produced by running `bioformats2raw` on a fileset that contains more than one image (series > 1):
series.ome.zarr # One converted fileset from bioformats2raw
├── .zgroup
├── .zattrs # Contains "bioformats2raw.layout" metadata
├── OME # Special group for containing OME metadata
│ ├── .zgroup
│ ├── .zattrs # Contains "series" metadata
│ └── METADATA.ome.xml # OME-XML file stored within the Zarr fileset
├── 0 # First image in the collection
├── 1 # Second image in the collection
└── ...
#### Attributes
(version0.4:bf2raw-attributes)=
The top-level `.zattrs` file must contain the `bioformats2raw.layout` key:
```{literalinclude} examples/bf2raw/image.json
language: json
```
If the top-level group represents a plate, the `bioformats2raw.layout` metadata will be present but
the "plate" key MUST also be present, takes precedence and parsing of such datasets should follow (plate metadata)(version0.4:plate-md). It is not
possible to mix collections of images with plates at present.
```{literalinclude} examples/bf2raw/plate.json
language: json
```
The `.zattrs` file within the OME group may contain the "series" key:
path: examples/ome/series-2.json
highlight: json
#### Details
(version0.4:bf2raw-details)=
Conforming groups:
- MUST have the value "3" for the "bioformats2raw.layout" key in their `.zattrs` metadata at the top of the hierarchy;
- SHOULD have OME metadata representing the entire collection of images in a file named "OME/METADATA.ome.xml" which:
- MUST adhere to the OME-XML specification but
- MUST use `` elements as opposed to ``, `` or ``;
- MAY make use of the [minimum specification](https://docs.openmicroscopy.org/ome-model/6.2.2/specifications/minimum.html).
Additionally, the logic for finding the Zarr group for each image follows the following logic:
- If "plate" metadata is present, images MUST be located at the defined location.
- Matching "series" metadata (as described next) SHOULD be provided for tools that are unaware of the "plate" specification.
- If the "OME" Zarr group exists, it:
- MAY contain a "series" attribute. If so:
- "series" MUST be a list of string objects, each of which is a path to an image group.
- The order of the paths MUST match the order of the "Image" elements in "OME/METADATA.ome.xml" if provided.
- If the "series" attribute does not exist and no "plate" is present:
- separate "multiscales" images MUST be stored in consecutively numbered groups starting from 0 (i.e. "0/", "1/", "2/", "3/", ...).
- Every "multiscales" group MUST represent exactly one OME-XML "Image" in the same order as either the series index or the group numbers.
Conforming readers:
- SHOULD make users aware of the presence of more than one image (i.e. SHOULD NOT default to only opening the first image);
- MAY use the "series" attribute in the "OME" group to determine a list of valid groups to display;
- MAY choose to show all images within the collection or offer the user a choice of images, as with HCS plates;
- MAY ignore other groups or arrays under the root of the hierarchy.
### "coordinateTransformations" metadata
(version0.4:trafo-md)=
"coordinateTransformations" describe a series of transformations that map between two coordinate spaces (defined by "axes").
For example, to map a discrete data space of an array to the corresponding physical space.
It is a list of dictionaries. Each entry describes a single transformation and MUST contain the field "type".
The value of "type" MUST be one of the elements of the `type` column in the table below.
Additional fields for the entry depend on "type" and are defined by the column `fields`.
| `identity` | | identity transformation, is the default transformation and is typically not explicitly defined
|
|---|
| `translation` | one of: `"translation":List[float]`, `"path":str` | translation vector, stored either as a list of floats (`"translation"`) or as binary data at a location in this container (`path`). The length of vector defines number of dimensions. |
|
|---|
| `scale` | one of: `"scale":List[float]`, `"path":str` | scale vector, stored either as a list of floats (`scale`) or as binary data at a location in this container (`path`). The length of vector defines number of dimensions. |
| type | fields | description
|
|---|
|
|---|
The transformations in the list are applied sequentially and in order.
### "multiscales" metadata
(version0.4:multiscale-md)=
Metadata about an image can be found under the "multiscales" key in the group-level metadata. Here, image refers to 2 to 5 dimensional data representing image or volumetric data with optional time or channel axes. It is stored in a multiple resolution representation.
"multiscales" contains a list of dictionaries where each entry describes a multiscale image.
Each "multiscales" dictionary MUST contain the field "axes", see (axes metadata)(#version0.4:axes-md).
The length of "axes" must be between 2 and 5 and MUST be equal to the dimensionality of the zarr arrays storing the image data (see "datasets:path").
The "axes" MUST contain 2 or 3 entries of "type:space" and MAY contain one additional entry of "type:time" and MAY contain one additional entry of "type:channel" or a null / custom type.
The order of the entries MUST correspond to the order of dimensions of the zarr arrays. In addition, the entries MUST be ordered by "type" where the "time" axis must come first (if present), followed by the "channel" or custom axis (if present) and the axes of type "space".
If there are three spatial axes where two correspond to the image plane ("yx") and images are stacked along the other (anisotropic) axis ("z"), the spatial axes SHOULD be ordered as "zyx".
Each "multiscales" dictionary MUST contain the field "datasets", which is a list of dictionaries describing the arrays storing the individual resolution levels.
Each dictionary in "datasets" MUST contain the field "path", whose value contains the path to the array for this resolution relative
to the current zarr group. The "path"s MUST be ordered from largest (i.e. highest resolution) to smallest.
Each "datasets" dictionary MUST have the same number of dimensions and MUST NOT have more than 5 dimensions. The number of dimensions and order MUST correspond to number and order of "axes".
Each dictionary in "datasets" MUST contain the field "coordinateTransformations", which contains a list of transformations that map the data coordinates to the physical coordinates (as specified by "axes") for this resolution level.
The transformations are defined according to (transformations metadata)(#version0.4:trafo-md). The transformation MUST only be of type `translation` or `scale`.
They MUST contain exactly one `scale` transformation that specifies the pixel size in physical units or time duration. If scaling information is not available or applicable for one of the axes, the value MUST express the scaling factor between the current resolution and the first resolution for the given axis, defaulting to 1.0 if there is no downsampling along the axis.
It MAY contain exactly one `translation` that specifies the offset from the origin in physical units. If `translation` is given it MUST be listed after `scale` to ensure that it is given in physical coordinates.
The length of the `scale` and `translation` array MUST be the same as the length of "axes".
The requirements (only `scale` and `translation`, restrictions on order) are in place to provide a simple mapping from data coordinates to physical coordinates while being compatible with the general transformation spec.
Each "multiscales" dictionary MAY contain the field "coordinateTransformations", describing transformations that are applied to all resolution levels in the same manner.
The transformations MUST follow the same rules about allowed types, order, etc. as in "datasets:coordinateTransformations" and are applied after them.
They can for example be used to specify the `scale` for a dimension that is the same for all resolutions.
Each "multiscales" dictionary SHOULD contain the field "name". It SHOULD contain the field "version", which indicates the version of the multiscale metadata of this image (current version is 0.4).
Each "multiscales" dictionary SHOULD contain the field "type", which gives the type of downscaling method used to generate the multiscale image pyramid.
It SHOULD contain the field "metadata", which contains a dictionary with additional information about the downscaling method.
```{literalinclude} examples/multiscales_strict/multiscales_example.json
language: json
```
If only one multiscale is provided, use it. Otherwise, the user can choose by
name, using the first multiscale as a fallback:
```python
datasets = []
for named in multiscales:
if named["name"] == "3D":
datasets = [x["path"] for x in named["datasets"]]
break
if not datasets:
# Use the first by default. Or perhaps choose based on chunk size.
datasets = [x["path"] for x in multiscales[0]["datasets"]]
```
### "omero" metadata (transitional)
(version0.4:omero-md)=
[=Transitional=] information specific to the channels of an image and how to render it
can be found under the "omero" key in the group-level metadata:
```json
"id": 1, # ID in OMERO
"name": "example.tif", # Name as shown in the UI
"version": "0.4", # Current version
"channels": [ # Array matching the c dimension size
{
"active": true,
"coefficient": 1,
"color": "0000FF",
"family": "linear",
"inverted": false,
"label": "LaminB1",
"window": {
"end": 1500,
"max": 65535,
"min": 0,
"start": 0
}
}
],
"rdefs": {
"defaultT": 0, # First timepoint to show the user
"defaultZ": 118, # First Z section to show the user
"model": "color" # "color" or "greyscale"
}
```
See https://docs.openmicroscopy.org/omero/5.6.1/developers/Web/WebGateway.html#imgdata
for more information.
The "omero" metadata is optional, but if present it MUST contain the field "channels", which is an array of dictionaries describing the channels of the image.
Each dictionary in "channels" MUST contain the field "color", which is a string of 6 hexadecimal digits specifying the color of the channel in RGB format.
Each dictionary in "channels" MUST contain the field "window", which is a dictionary describing the windowing of the channel.
The field "window" MUST contain the fields "min" and "max", which are the minimum and maximum values of the window, respectively.
It MUST also contain the fields "start" and "end", which are the start and end values of the window, respectively.
### "labels" metadata
(version0.4:labels-md)=
The special group "labels" found under an image Zarr contains the key `labels` containing
the paths to label objects which can be found underneath the group:
```json
{
"labels": [
"orphaned/0"
]
}
```
Unlisted groups MAY be labels.
### "image-label" metadata
(version0.4:label-md)=
Groups containing the `image-label` dictionary represent an image segmentation
in which each unique pixel value represents a separate segmented object.
`image-label` groups MUST also contain `multiscales` metadata and the two
"datasets" series MUST have the same number of entries.
The `image-label` dictionary SHOULD contain a `colors` key whose value MUST be a
list of JSON objects describing the unique label values. Each color object MUST
contain the `label-value` key whose value MUST be an integer specifying the
pixel value for that label. It MAY contain an `rgba` key whose value MUST be an array
of four integers between 0 and 255 `[uint8, uint8, uint8, uint8]` specifying the label
color as RGBA. All the values under the `label-value` key MUST be unique. Clients
who choose to not throw an error SHOULD ignore all except the _last_ entry.
Some implementations MAY represent overlapping labels by using a specially assigned
value, for example the highest integer available in the pixel range.
The `image-label` dictionary MAY contain a `properties` key whose value MUST be a
list of JSON objects which also describes the unique label values. Each property object
MUST contain the `label-value` key whose value MUST be an integer specifying the pixel
value for that label. Additionally, an arbitrary number of key-value pairs
MAY be present for each label value denoting associated metadata. Not all label
values must share the same key-value pairs within the properties list.
The `image-label` dictionary MAY contain a `source` key whose value MUST be a JSON
object containing information on the image the label is associated with. If included,
it MAY include a key `image` whose value MUST be a string specifying the relative
path to a Zarr image group. The default value is "../../" since most labels are stored
under a subgroup named "labels/" (see above).
The `image-label` dictionary SHOULD contain a `version` key whose value MUST be a string
specifying the version of the image-label specification.
```{literalinclude} examples/label_strict/colors_properties.json
language: json
```
### "plate" metadata
(version0.4:plate-md)=
For high-content screening datasets, the plate layout can be found under the
custom attributes of the plate group under the `plate` key in the group-level metadata.
The `plate` dictionary MAY contain an `acquisitions` key whose value MUST be a list of
JSON objects defining the acquisitions for a given plate to which wells can refer to. Each
acquisition object MUST contain an `id` key whose value MUST be an unique integer identifier
greater than or equal to 0 within the context of the plate to which fields of view can refer
to (see [well metadata](version0.4:well-md)).
Each acquisition object SHOULD contain a `name` key whose value MUST be a string identifying
the name of the acquisition. Each acquisition object SHOULD contain a `maximumfieldcount`
key whose value MUST be a positive integer indicating the maximum number of fields of view for the
acquisition. Each acquisition object MAY contain a `description` key whose value MUST be a
string specifying a description for the acquisition. Each acquisition object MAY contain
a `starttime` and/or `endtime` key whose values MUST be integer epoch timestamps specifying
the start and/or end timestamp of the acquisition.
The `plate` dictionary MUST contain a `columns` key whose value MUST be a list of JSON objects
defining the columns of the plate. Each column object defines the properties of
the column at the index of the object in the list. Each column in the physical plate
MUST be defined, even if no wells in the column are defined. Each column object MUST
contain a `name` key whose value is a string specifying the column name. The `name` MUST
contain only alphanumeric characters, MUST be case-sensitive, and MUST NOT be a duplicate of any
other `name` in the `columns` list. Care SHOULD be taken to avoid collisions on
case-insensitive filesystems (e.g. avoid using both `Aa` and `aA`).
The `plate` dictionary SHOULD contain a `field_count` key whose value MUST be a positive integer
defining the maximum number of fields per view across all wells.
The `plate` dictionary SHOULD contain a `name` key whose value MUST be a string defining the
name of the plate.
The `plate` dictionary MUST contain a `rows` key whose value MUST be a list of JSON objects
defining the rows of the plate. Each row object defines the properties of
the row at the index of the object in the list. Each row in the physical plate
MUST be defined, even if no wells in the row are defined. Each defined row MUST
contain a `name` key whose value MUST be a string defining the row name. The `name` MUST
contain only alphanumeric characters, MUST be case-sensitive, and MUST NOT be a duplicate of any
other `name` in the `rows` list. Care SHOULD be taken to avoid collisions on
case-insensitive filesystems (e.g. avoid using both `Aa` and `aA`).
The `plate` dictionary SHOULD contain a `version` key whose value MUST be a string specifying the
version of the plate specification.
The `plate` dictionary MUST contain a `wells` key whose value MUST be a list of JSON objects
defining the wells of the plate. Each well object MUST contain a `path` key whose value MUST
be a string specifying the path to the well subgroup. The `path` MUST consist of a `name` in
the `rows` list, a file separator (`/`), and a `name` from the `columns` list, in that order.
The `path` MUST NOT contain additional leading or trailing directories.
Each well object MUST contain both a `rowIndex` key whose value MUST be an integer identifying
the index into the `rows` list and a `columnIndex` key whose value MUST be an integer identifying
the index into the `columns` list. `rowIndex` and `columnIndex` MUST be 0-based. The
`rowIndex`, `columnIndex`, and `path` MUST all refer to the same row/column pair.
For example the following JSON object defines a plate with two acquisitions and
6 wells (2 rows and 3 columns), containing up to 2 fields of view per acquisition.
```{literalinclude} examples/plate_strict/plate_6wells.json
:language: json
```
The following JSON object defines a sparse plate with one acquisition and
2 wells in a 96 well plate, containing one field of view per acquisition.
```{literalinclude} examples/plate_strict/plate_2wells.json
:language: json
```
### "well" metadata
(version0.4:well-md)=
For high-content screening datasets, the metadata about all fields of views
under a given well can be found under the "well" key in the attributes of the
well group.
The `well` dictionary MUST contain an `images` key whose value MUST be a list of JSON objects
specifying all fields of views for a given well. Each image object MUST contain a
`path` key whose value MUST be a string specifying the path to the field of view. The `path`
MUST contain only alphanumeric characters, MUST be case-sensitive, and MUST NOT be a duplicate
of any other `path` in the `images` list. If multiple acquisitions were performed in the plate,
it MUST contain an `acquisition` key whose value MUST be an integer identifying the acquisition
which MUST match one of the acquisition JSON objects defined in the plate metadata (see [plate-metadata](version0.4:plate-md)).
The `well` dictionary SHOULD contain a `version` key whose value MUST be a string specifying the
version of the well specification.
For example the following JSON object defines a well with four fields of
view. The first two fields of view were part of the first acquisition while
the last two fields of view were part of the second acquisition.
```{literalinclude} examples/well_strict/well_4fields.json
:language: json
```
The following JSON object defines a well with two fields of view in a plate with
four acquisitions. The first field is part of the first acquisition, and the second
field is part of the last acquisition.
```{literalinclude} examples/well_strict/well_2fields.json
:language: json
```
## Specification naming style
(version0.4:naming-style)=
Multi-word keys in this specification should use the `camelCase` style.
NB: some parts of the specification don't obey this convention as they
were added before this was adopted, but they should be updated in due course.
## Implementations
(version0.4:implementations)=
Projects which support reading and/or writing OME-NGFF data include:
- [bigdataviewer-ome-zarr](https://github.com/mobie/bigdataviewer-ome-zarr)
- Fiji-plugin for reading OME-Zarr.
- [bioformats2raw](https://github.com/glencoesoftware/bioformats2raw)
- A performant, Bio-Formats image file format converter.
- [omero-ms-zarr](https://github.com/ome/omero-ms-zarr)
- A microservice for OMERO.server that converts images stored in OMERO to OME-Zarr files on the fly, served via a web API.
- [idr-zarr-tools](https://github.com/IDR/idr-zarr-tools)
- A full workflow demonstrating the conversion of IDR images to OME-Zarr images on S3.
- [OMERO CLI Zarr plugin](https://github.com/ome/omero-cli-zarr)
- An OMERO CLI plugin that converts images stored in OMERO.server into a local Zarr file.
- [ome-zarr-py](https://github.com/ome/ome-zarr-py)
- A napari plugin for reading ome-zarr files.
- [vizarr](https://github.com/hms-dbmi/vizarr/)
- A minimal, purely client-side program for viewing Zarr-based images with Viv & ImJoy.
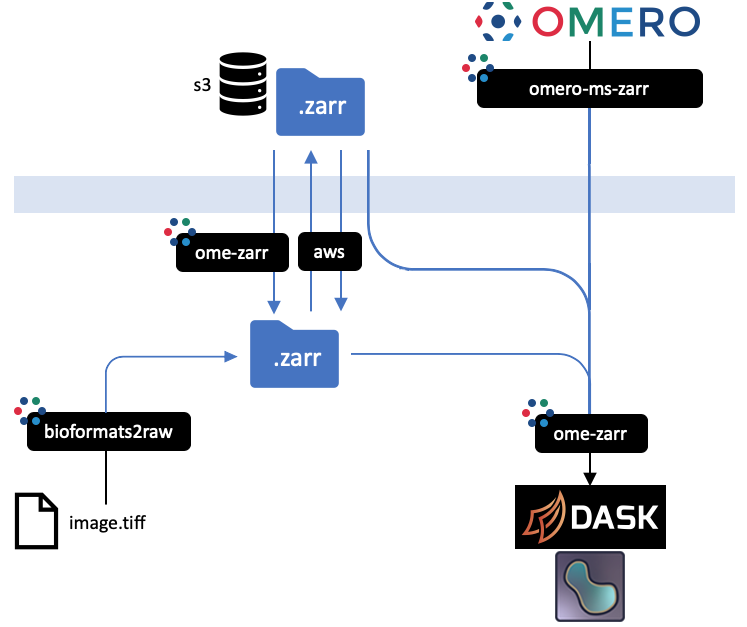 All implementations prevent an equivalent representation of a dataset which can be downloaded or uploaded freely. An interactive
version of this diagram is available from the [OME2020 Workshop](https://downloads.openmicroscopy.org/presentations/2020/Dundee/Workshops/NGFF/zarr_diagram/).
Mouseover the blackboxes representing the implementations above to get a quick tip on how to use them.
Note: If you would like to see your project listed, please open an issue or PR on the [ome/ngff](https://github.com/ome/ngff) repository.
## Other resources
```{toctree}
:maxdepth: 1
examples/index
schemas/index
```
All implementations prevent an equivalent representation of a dataset which can be downloaded or uploaded freely. An interactive
version of this diagram is available from the [OME2020 Workshop](https://downloads.openmicroscopy.org/presentations/2020/Dundee/Workshops/NGFF/zarr_diagram/).
Mouseover the blackboxes representing the implementations above to get a quick tip on how to use them.
Note: If you would like to see your project listed, please open an issue or PR on the [ome/ngff](https://github.com/ome/ngff) repository.
## Other resources
```{toctree}
:maxdepth: 1
examples/index
schemas/index
```  All implementations prevent an equivalent representation of a dataset which can be downloaded or uploaded freely. An interactive
version of this diagram is available from the [OME2020 Workshop](https://downloads.openmicroscopy.org/presentations/2020/Dundee/Workshops/NGFF/zarr_diagram/).
Mouseover the blackboxes representing the implementations above to get a quick tip on how to use them.
Note: If you would like to see your project listed, please open an issue or PR on the [ome/ngff](https://github.com/ome/ngff) repository.
## Other resources
```{toctree}
:maxdepth: 1
examples/index
schemas/index
```
All implementations prevent an equivalent representation of a dataset which can be downloaded or uploaded freely. An interactive
version of this diagram is available from the [OME2020 Workshop](https://downloads.openmicroscopy.org/presentations/2020/Dundee/Workshops/NGFF/zarr_diagram/).
Mouseover the blackboxes representing the implementations above to get a quick tip on how to use them.
Note: If you would like to see your project listed, please open an issue or PR on the [ome/ngff](https://github.com/ome/ngff) repository.
## Other resources
```{toctree}
:maxdepth: 1
examples/index
schemas/index
```